INTRODUCTION
MicroRNAs (miRNAs) are a group of short noncoding RNA molecules usually having a length of about 22 nucleotides (O’Brien et al., 2018). miRNAs are basically endogenous biomolecules and most of them have remained conserved throughout the course of evolution, which suggests that they have important biological functions. The regulatory role of the miRNA has been discussed in several studies (Bartel, 2004; Lau et al., 2001; Reinhart et al., 2000). They were first discovered in the nematode, Caenorhabditis elegans, produced by the gene known as lin-4 and were found to post-transcriptionally downregulate the lin-14mRNA (Lee et al., 1993; Wightman et al., 1993). Later miRNAs were also discovered in eukaryotes including animals, plants, human beings, and some viruses. Various scientific works have revealed that miRNAs are mainly involved in gene silencing and in the post-transcriptional regulation of protein expression (de Carvalho et al., 1992; Napoli et al., 1990). In addition to this, they also play an important role in several biological processes including the regulation of cell cycle, cell differentiation, leaf and flower development, fat metabolism, immunity, and cell death (Hannon, 2002; Romano and Macino, 1992).
miRNAs have specific target mRNAs with which they bind by complementary base pairing to silence that mRNA (Catalanotto et al., 2016). However, the manner in which they function in plant and animal is slightly different. In animals, two Rnase III-type proteins – Drosha (in nucleus) and Dicer (in cytoplasm) synthesize the mature miRNA in two stages from primary miRNAs or pre-miRNAs; whereas in plants, a single RNase III enzyme named Dicer-like 1 (DCL1) is responsible for the synthesis of mature miRNAs from pre-miRNAs and the whole process takes place entirely in the nucleus (Kim, 2005). Argonaute (Ago) subfamily proteins, then bind to these newly formed matured miRNAs. After this, specific mRNAs are targeted by these processed miRNAs to carry out their regulatory functions.
Mangroves comprise a group of highly specialized plant community, which have the capability to thrive under constant physiological stress. Tropical and subtropical river deltas, creeks are home to majority of the mangrove vegetation across globe (Tomlinson, 1986). They act as the primary defense line against the various typhoons and sea storms. The mangrove ecosystem is also regarded as immensely economically viable ecosystem. A report estimated the economic value to be worth about US$186 million per year (FAO, 2007). Sundarbans mangrove forest situated at the Indian sub-continent is the most diverse among all the mangrove forests across globe, with the maximum species diversity (Das, 1999). (Naskar et al., 1987) reported 36 true mangroves, 28 associate mangrove, and 7 obligatory mangrove species which belong to 29 families and 49 genera. However, according to recent reports, growth of some of the species has become stunted and some are on the verge of extinction (IUCN, 2020; Upadhyay et al., 2002). Several natural factors, such as global warming, sea level rise, increment in soil and water salinity in the mangrove estuaries, have caused havoc damage to these population (Dasgupta et al., 2010; Nandy et al., 2007). These factors have added up to the stress factors that mangrove vegetation already had to cope up due to their habitat; high salinity, periodic inundation, and high UV radiation being some of them. It has been inferred from a number of studies that any kind of stress, be it biotic and/or abiotic, increases the rate of reactive oxygen radical production in the plants (Asada, 1994; Prochazkova and Wilhelmova, 2007). Mangroves being constantly under severe physiological stress constantly generates huge amount of these free radicals, which, in turn, has the capacity of oxidizing proteins, DNA, peroxidation of lipid, and ultimately breakdown of cells (Twigg et al., 1998). Plants, on the other hand, have a defense mechanism to scavenge off these excess ROS to maintain homeostasis inside the plant-system, comprising several antioxidant and hydrolyzing enzymes, numerous secondary metabolites, some osmolytes, etc. (Dasgupta et al., 2014; Demiral et al., 2005; Parida et al., 2004) However, recent studies have elucidated the role of miRNA in regulating the homeostasis in plants (Ding et al., 2020; Gu et al., 2017; Zhu et al., 2020). Although some studies have been conducted on the ROS scavenging abilities of some mangrove species, study on miRNA with respect to their role in stress management is very rare in Sundarban perspective (Dasgupta et al., 2017). In this present work, we have studied the miRNAs present in the mangrove vegetation and how they effect in maintaining homeostasis inside the plant, if any such miRNA is found. Among all the mangrove flora present, Rhizophora species has been selected for this study.
MATERIALS AND METHODS
Retrieval of Data and Trimming
We started our study with three Rhizophora species, that is, Rhizophora mucronata, Rhizophora stylosa, and Rhizophora apiculata. However, due to non-availability of significant amount of data, we continued the study only with R. apiculata. miRNA database named miRBase was used to retrieve the miRNA sequences of available 10615 plant sequences (Kozomara et al., 2019). PRINSEQ was then used to remove the redundant sequences (Schmieder and Edwards, 2011). Then, the R. apiculata EST database of National Centre for Biotechnology Information (NCBI) was used to search the homologs for the remaining 5889 miRNA sequences that were left after the redundancy check.
Potential miRNAs Identification
Basic Local Alignment Search Tool (BLAST), a NCBI tool, was used for the alignment of sequences against the publicly available database (Altschul et al., 1990). This tool was used for the alignment of all the above-mentioned 5889 miRNA sequences against the R. apiculata EST database. The nucleotide match size between query and database was set to 15 with expectation value 0.01. The match sequence less than this value were not taken into account. The selected ESTs, after the redundancy check, were subjected to BLASTx to eliminate the protein-coding sequences. BLAST was performed again against the NCBI nucleotide database to exclude the other form of RNAs such as rRNA, tRNA, snRNA, or snoRNA.
RNA Secondary Structure Prediction
The mfold Web Server was used for obtaining the secondary structures of the selected candidates (Zuker, 2003). The parameters were kept as default. The potential miRNAs were identified based on the following criteria – (i) This screening was done by keeping the position of the miRNA on the hairpin, (ii) the minimum number of residues in miRNA as 15, (iii) the maximum number of unpaired residues as 6, (iv) the maximum number of G-U pairs in miRNA as 5, (v) the maximum size for a bulge in miRNA sequence as 5 nt, (vi) the negative minimal folding free energy (MFE) was considered low, and (vii) the minimal folding free energy index (MFEI) was considered to be high. MFEI is equal to (MFE/length of the RNA sequence*100)/(G+C) %.
Potential miRNA Target Prediction
The predicted miRNAs from R. apiculata were then analyzed for obtaining the potential targets using psRNATarget web server (Dai and Zhao, 2011). The database of Arabidopsis thaliana was used for searching potential targets, as no miRNA from R. apiculata was reported till the time of this study.
RESULTS AND DISCUSSION
All the previously reported plant miRNA were downloaded from the miRNA database. In this study, all the 10615 previously reported mature miRNA sequences were used to find their homologs in the mangrove species, R. apiculata. This step is important as till date no study on the miRNA on R. apiculata has been reported, so in silico screening of their homologs from the already reported species is an excellent way for predicting the miRNAs which might be present in the species under consideration. According to various studies, miRNAs are said to play important role in plant growth and development (Gong et al., 2015; Singh et al., 2018). They are also related with the stress responses in plants (Khraiwesh et al., 2012; Sunkar et al., 2012). Hence, in this study, we have predicted potential miRNAs, using the expressed sequence tag database of R. apiculata.
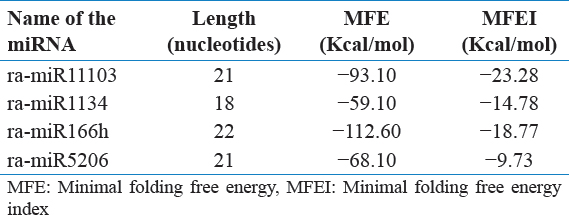
The next step, that is, redundancy check eliminated the duplicate entries of the miRNAs which were collected from the database. In the quest of finding the nucleotide sequences of R. apiculata which might contain the miRNAs, the 5889 unique sequences were used as query to find their homologs present in R. apiculata EST database. Nucleotide BLAST was performed to align the ESTs with the query miRNA sequences. Further, BLASTx was performed with the shortlisted ESTs to eliminate the protein coding sequences and non-coding RNAs. The resulting sequences thus found have the potential to be or to contain miRNAs. Next step was predicting the secondary structure of RNA using mfold software. For a RNA or nucleotide strand to act as miRNAs they have to mandatorily be in a specific conformation, or else their activity would be absent. The nucleotides that were homologous to the miRNAs were selected and inspected for this conformation by taking into account the seven criterions mentioned earlier. Among all the ESTs taken under consideration the criterion got matched for four ESTs. They were named as ra-miR11103, ra-miR1134, ra-miR166h, and ra-miR5206 [Table 1]. The Minimal Folding Free Energy (MFE) for ra-miR1134 was found to be −59.1 Kcal/mol, whereas the minimal folding free energy index (MFEI) for the same was calculated to be −14.78 Kcal/mol [Table 1].
Table 1: Predicted miRNA molecules
The role of miRNA in helping plants to sustain in stressed conditions and grow normally has been mentioned (Noman and Aqeel, 2017). To find out the role of these newly predicted miRNAs in helping the mangrove species in such hostile condition, we searched for the probable target genes for the 4 miRNAs. psRNA webserver was used for this study. Due to the non-availability of cDNA library for R. apiculata, the cDNA library for A. thaliana was utilized for this study due to the fact that being a model organism A. thaliana has been extensively studied (Allen et al., 2004; Fahlgren et al., 2010; Liu et al., 2008). The result generated from this study was evaluated for understanding the role of these miRNAs in maintaining cellular homeostasis. The miRNA ra-miR11103 generated a total of 183 hits, which basically shows the versatility of target genes for this miRNA. On further analysis of this hits, it was found that the target gene for miRNA 11103 includes DNAJ heat shock protein, RNA directed DNA polymerase, glutathione S-transferase (GST), CBL interacting protein kinase, chlorophyll a-b binding protein, cytokinin receptor CRE1b, histidine kinase, serine/threonine protein kinase, Mannose-1-phosphate guanylyltransferase, and NADH:ubiquinone oxidoreductase among others. Eventually all of the proteins or enzymes mentioned have significant role to play in different stages of growth of the plant. Zhichang et al., 2010, reported the specific role of the DNAJ heat shock protein 70 in stress tolerance of plants. Over expression of this gene has been directly correlated with NaCl tolerance in plants. The pivotal role of GST in combating abiotic stress has been discussed by various scientists (Hossain et al., 2006; Kumar and Trivedi, 2018). Chlorophyll a-b binding protein does play a significant role as a signaling molecule of stress (Costa et al., 2008). A signaling molecule like this sends signal inside the plant system regarding the presence and absence and also intensity of the stress and accordingly the plant starts to produce the various secondary metabolites, enzymes, etc., to combat against the stress. Mannose-1-phosphate guanylyltransferase is an enzyme belonging to transferases family. Recent study has revealed that Mannose-1-phosphate guanylyltransferase is a key component which gives plants resistance against salinity stress (Kumar et al., 2012). First isolated from a salt tolerant rice variety Pokkali, this enzyme enables yeasts to grow at salinity as high as 1.2 M.
The study on ra-miR1134 showed a total of 145 hits. Some of the predicted genes for this miRNA included RING finger family proteins, disease resistance Cf-2 like protein, sucrose transport protein SUC4, serine carboxypeptidase, actin interacting protein, tobacco mosaic virus (TMV) resistance protein, protein kinase family protein, and TATA binding protein. The role of RING finger family proteins in regulation of stress responses as well as in disease susceptibility has been reported by Hong et al., 2007. Sugar molecules are produced in large quantity during osmotic stressed condition in plants to increase the cell sap water potential. This phenomenon makes the water potential inside the cell sap more negative, which helps the plant to withstand high osmotic stress. The role of Sucrose transport protein SUC4 as a signaling molecule during abiotic stress has been observed by various scientists (Durand et al., 2016; Eom et al., 2015). TATA box binding protein is a well-known class of proteins which regulate the DNA replication of any living organism, as attachment of the TATA box with the TATA box binding protein itself initiates the DNA replication process. Disease resistance cf-2 like protein, TMV resistance protein these are a classic example of proteins which confers resistance to plants against certain pathogenic organisms. TMV resistance protein confers resistance to plants against pathogen such as Cladosporium fulvum, which is a leaf mold fungus (Dixon et al., 1996) and TMV is a widespread disease which affect a number a crops across the globe. Hence, as predicted by the software one of the major roles of this miRNA might be regulation of the disease resistance or abiotic stress tolerance of course along with other roles to maintain homeostasis inside the plant system; which is in contrast with the previously described miRNA where the major role was regulating abiotic stress responses. The other two miRNAs – ra-miR166h and ra-miR5206 generated 122 and 148 responses, respectively. All the target sequences for these two, as achieved by this study also points out toward stress tolerance, be it biotic or abiotic. These findings are actually at par with the fact that the mangrove plants constantly thrive under immense physiological stress and to maintain homeostasis for proper growth, in such hostile environment they need very strong defense mechanism (Asaeda and Barnuevo, 2019; Dasgupta et al., 2011).
CONCLUSION
The salinity in the Sundarbans estuary in on a constant rise since past few decades (Nandy et al., 2007) and the result of that has become detrimental for various mangrove species such as Heritiera fomes, Xylocarpus granatum, and Nypa fruticans (IUCN, 2020). These plants are almost on the verge of extinction and in the present rate some will get extinct within a decade or so. Restoration program for such species is being taken up in some parts and their conservation cannot be done properly unless enough genetic data are available. However, genetic information available for Sundarbans mangroves is too sparse. Hence, the first step towards the restoration programs should include in depth genetic analysis for these species, which will generate the proper reason for their gradual extinction. In the present study, it was found out that these miRNAs present in R. apiculata, which is a profusely growing species in Indian Sundarbans, target many genes which are the backbone of growth and regulation of plants. Hence, further experimental validation of the predictions received from this analysis using the plant samples from Indian Sundarbans mangrove forest might throw light towards the genetic composition of the mangroves, and that data being compared with those species which are now in distressed condition, might throw light towards the reason for their gradual disappearance, which again, in turn, might be helpful in creating strategies for the proper restoration of those species.
Financial Support and Sponsorship
Nil.
Conflicts of Interest
There are no conflicts of interest.